A recent study undertaken by an international team of academics and published in the academic journal PLOS Pathogens, has investigated the causes that turn localised infections into the life-threatening systematic disease sepsis, and has found genetic evidence pointing at one single bacteria as the cause.
This is the first time that such as discovery has been made through examination, in mice, of the events leading to sepsis by Streptococcus pneumoniae (pneumococcus), a major human pathogen.
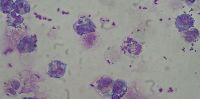
Professor Marco Oggioni from the University of Leicester's Department of Genetics has, together with his team, found that the bacteria causing sepsis was started by a single pneumococcal cell in most cases.
As an interdisciplinary collaboration between the Departments of Genetics, Infection Immunity and Inflammation and Mathematics at the University of Leicester, Professor Richard Moxon at the University of Oxford, and scientists from overseas including the University of Siena, this study was groundbreaking.
Professor Oggioni explained how data in experimental infection models indicated that strategies were needed both for targeting bacteria once the infection has progressed, and for early intervention schemes preventing the one-single cell that starts the disease process. He added that the research had for the first time provided genetic evidence for a single cell origin of bacterial invasive infection. While this scenario was hypothesised over five decades ago, it had only been phenotypic, whereas now statistical evidence could be obtained for this event.
Under normal circumstances, when different bacteria are used in models of experimental infection of hosts who have not previously encountered the same pathogen, the vast majority is destroyed rapidly by the host's innate immune system. In the researcher's model, a dose of one million bacteria is needed to induce systemic disease in about half of the hosts in the study.
This is in stark contrast to a much lower number of bacteria thought to make up the starting "seed" that leads to the development of systemic infection, and the assumption is that there must be at least one "bottlenecks" in the development of the disease.
In order to investigate these bottlenecks, the team of researchers injected mice with a combination of three different variants of S. pneumoniae. Roughly half the mice developed sepsis and in almost all of those cases the bacteria causing sepsis were derived from only one of the three variants used in the initial challenge.
Through the use of statistical analysis, as well as direct DNA sequencing, the researchers were able to demonstrate that the bacterial population causing sepsis was started by a single pneumococcal cell in most of the cases.
While analysing what causes the immune system to resist most of the injected bacteria, it was discovered that macrophages in the spleen, a type of immune cell capable of gobbling up bacteria, are the main contributors to an efficient immune response to this pathogen.
The team’s findings suggest that if bacteria survive this initial hurdle, a single 'founder' bacterium multiplies and re-enters the bloodstream, where its descendants come under strong selective pressure that dynamically shapes the subsequent bacterial population. This condition results in the sepsis. Data has also uncovered no obvious characteristics in the single bacterium leading to sepsis, suggesting that it is random events determining which of the injected bacteria survives and multiplies to cause disease.
This discovery could also potentially apply to human systemic infections and prove vital to understanding sepsis, as the causes of the disease are still largely unknown to the scientific community.
Dr Oggioni believes that this newfound knowledge provides an important paradigm and essential information which, the team hopes, will lead to changes in therapeutic protocols in order to positively impact success of treatment outcome.
Source: Science Daily
24 March 2014
Latest Articles
Research, Sepsis, bacteria, blood cells, Sepsis Therapy, immune system
A recent study undertaken by an international team of academics and published in the academic journal PLOS Pathogens, has investigated the causes that turn...